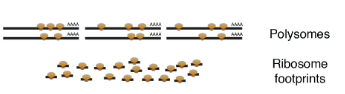
Ribosome profiling is a sequencing tecnique that detects regions in mRNAs that are being translated. Using this technique, researchers have observed mysterious patterns of translation in many transcripts believed to be non-coding (lncRNAs, or long non-coding RNAs). The patterns are very similar to those observed in protein-coding genes but the translated proteins are generally smaller. Aside from their sequence, we know nothing about these peptides. Are they functional? Do they reflect some background noise of the translation machinery?
In a recent study published in bioRxiv we have investigated the signatures of selection in proteins translated from lncRNAs, using phylogenetic conservation and single nucleotide polymorphism (SNP) data. We have found that hundreds of mouse lncRNAs produce short functional proteins and thus should be considered protein coding genes. However, the largest part of translated lncRNAs appears to correspond to non-functional peptides. We conclude that, translation, like transcription, is pervasive. Due to this activity many peptides can be tested for new functions, facilitating the birth of new genes de novo.
This preprint was selected by the NODE (July 2016). It has also appeared at redcedar PRBB blog. The work was presented at XXI Evolution and Population Genetics Seminar Oct 3-5 2016 Sitges (Barcelona).


Pingback: Pervasive translation of lncRNAS | redcedar